eGoT: Enhanced Graph-of-Thoughts for Multi-Hop Knowledge Retrieval and Hypothesis Generation in Biomedicine
A retrieval-augmented generation algorithm that builds knowledge graphs directly from biomedical literature and reasons over them with an iterative graph-of-thoughts process. Accepted at ISMB 2026, to appear in Bioinformatics.

The lab’s paper “eGoT: Enhanced Graph-of-Thoughts for Multi-Hop Knowledge Retrieval and Hypothesis Generation in Biomedicine” has been accepted at ISMB 2026 and will appear in Bioinformatics. The work, led by Nihar Sanda with contributions from Benjamin M. Gyori, Vito Quaranta, Auroop Ganguly, and Ayan Paul, introduces eGoT, a retrieval-augmented generation algorithm that builds a knowledge graph directly from a corpus of biomedical literature and answers natural-language questions through an iterative graph-of-thoughts reasoning process.
This post provides an accessible summary of the approach for a general research audience, followed by a more detailed technical description of the method, its design choices, and its empirical behavior across benchmarks and biomedical case studies.
A Semi-Technical Summary
Biomedical knowledge is fragmented. Mechanistic insights, molecular data, and clinical observations relevant to a single question are routinely scattered across thousands of publications. PubMed alone catalogs roughly thirty-nine million articles. Answering even a moderately complex biomedical question often requires a researcher to piece together evidence from disparate sources and trace multi-hop reasoning chains, for example from a genetic variant to a downstream phenotype, or from an environmental exposure to an autoimmune manifestation.
Large language models can help, but on their own they hallucinate, lack provenance, and become outdated. Retrieval-augmented generation (RAG) addresses these issues by grounding model outputs in retrieved evidence. Vector-based RAG works well for shallow open-domain questions, but it struggles to chain together evidence across multiple sources when answering biomedical questions that span genes, mechanisms, environments, and clinical outcomes. Graph-based RAG approaches improve on this by reasoning over structured knowledge graphs, but most existing systems either depend on pre-curated knowledge bases or use single-pass graph traversal that limits reasoning to one or two hops.
eGoT addresses these limitations with two components. First, an end-to-end pipeline constructs a knowledge graph directly from unstructured publications, including entity-relation extraction, entity standardization, and rule-based, lexical, and community-based relationship inference. Second, a retrieval algorithm based on the graph-of-thoughts paradigm iteratively generates exploratory thoughts about a query, scores them, retrieves graph-grounded evidence for the highest-scoring ones, and then generates new thoughts conditioned on the accumulated evidence. The result is an answer that draws on multi-hop reasoning chains traversed across the graph, with every claim traceable back to specific source documents.
On standard RAG benchmarks (Ultradomain Legal, Ultradomain Agriculture, Ultradomain Mix, HotpotQA, and MultiHopRAG) eGoT outperforms a wide range of state-of-the-art retrieval methods, including HiRAG, LightRAG, NaiveRAG, GraphRAG, KAG, HippoRAG, HopRAG, SiReRAG, PathRAG, HyperGraphRAG, and TH-RAG. Two biomedical case studies, one on small cell lung cancer (SCLC) and one on cross-domain integration of lupus and UV radiation, demonstrate the system’s ability to support expert-curated question answering and generate testable hypotheses across traditionally siloed domains.
Why Multi-Hop Reasoning Over Biomedical Literature Matters
The notion of “undiscovered public knowledge”, introduced by Don Swanson in 1986, captures a long-standing intuition in biomedical research: many of the most important connections between genes, mechanisms, and diseases are already implicit in the literature, but spread across publications that no single researcher can read in full. Computational methods that can reason over and synthesize this distributed knowledge have the potential to accelerate discovery by surfacing latent connections that would otherwise require an extensive manual review.
Recent biomedical RAG systems have made progress on this problem. KRAGEN combines knowledge graphs with graph-of-thoughts prompting for biomedical problem solving. MedRAG constructs hierarchical diagnostic knowledge graphs for clinical decision support. i-MedRAG enables iterative follow-up queries to refine retrieval. Each of these systems contributes important pieces of the puzzle, but they tend to target clinical diagnosis or assume a pre-existing curated knowledge base, and their reasoning typically extends only one or two hops from the entities mentioned in the query. Many of the most consequential biomedical questions require deeper, cross-domain reasoning.
eGoT pushes on both fronts. It constructs the underlying knowledge graph from primary literature on demand, so the system is not limited to whatever ontology a curator has already produced. And it replaces single-pass graph traversal with an iterative reasoning loop, in which an LLM proposes new thoughts at each iteration based on the evidence retrieved so far, enabling the discovery of multi-hop chains of arbitrary depth.
The eGoT Pipeline
The end-to-end eGoT pipeline has two stages: knowledge graph construction from a document corpus, and graph-of-thoughts retrieval over the constructed graph. Both stages are LLM-driven, and the choice of LLM is configurable. The reported experiments use LLaMA-4 Scout (109B parameters) for graph construction and either DeepSeek-V3 (671B parameters) or GPT-4o for retrieval and answer generation, with sentence-transformers/all-MiniLM-L6-v2 for chunk and entity embeddings.
Knowledge Graph Construction
The graph is built in three stages. First, each document is partitioned into overlapping text chunks with a shared context window, and an LLM is prompted to extract subject-predicate-object triples from each chunk. Predicates are constrained to be at most three words to keep them concise and improve graph connectivity. The resulting initial graph G0 is the union of all extracted triples, with each triple linked back to its source chunk and parent document.
Second, raw entity mentions are standardized. Entities such as “Systemic Lupus Erythematosus”, “Lupus”, and “SLE” routinely appear in different surface forms across the literature, and treating them as distinct nodes fragments the graph. The standardization step normalizes entity strings, clusters near-duplicates by lexical and semantic similarity, and selects a canonical representative per cluster, weighting candidates by frequency and length.
Third, hidden relationships are inferred to enrich the graph. Transitive inference adds edges of the form (h, rinferred, t) when intermediate paths exist. Lexical inference adds edges between entities that share roots or morphological variants. Community-based inference identifies disconnected components and uses an LLM to propose plausible cross-community relationships, with the LLM constrained to short predicates and grounded in retrieved triples from each community.
Finally, the graph is organized hierarchically. Each triple is linked to its source chunk, and each chunk is linked to its parent document. Vector embeddings are computed and stored for every chunk, supporting efficient semantic search at query time while preserving document-level provenance.
Iterative Thought Generation and Evaluation
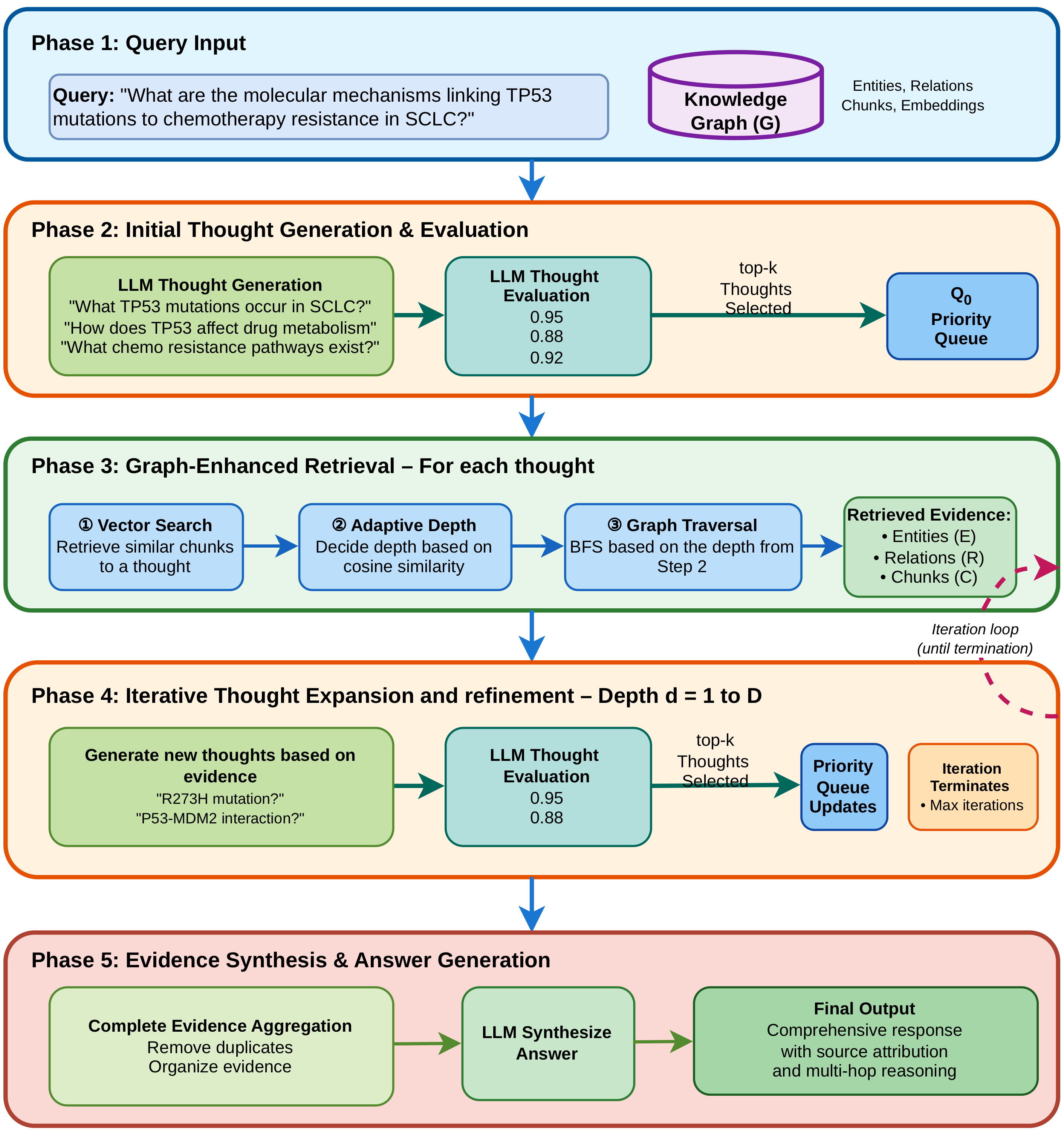
Given a query, eGoT does not retrieve directly. Instead, it first decomposes the query into a small set of exploratory thoughts through an LLM call, where each thought represents a distinct reasoning direction or hypothesis that could contribute to answering the question. The paper uses two to four thoughts per iteration to balance breadth of exploration against compute cost.
A separate LLM call then scores each thought on its potential contribution to answering the original query, producing a score in the unit interval. Thoughts are pushed onto a priority queue and explored in order of score, so that compute is concentrated on the most promising reasoning paths. Removing this evaluation step, the paper reports, degrades context precision sharply, since the algorithm then explores random thoughts.
The reasoning loop proceeds for up to a maximum depth (typically two iterations in the benchmarks, three or more for larger graphs). At each depth, the top-k thoughts from the queue are processed: each thought triggers a graph-enhanced retrieval, the retrieved evidence is summarized, and the cumulative evidence state is used as context for generating new thoughts at the next depth. The loop terminates when the maximum depth is reached, the priority queue is exhausted, or no novel thoughts are produced.
Adaptive Graph-Enhanced Retrieval
The retrieval step is the second technical novelty in the paper. For each thought, eGoT first performs a vector similarity search over chunk embeddings to identify candidate chunks. The entities present in those chunks become starting points for graph traversal. For each starting entity, the depth of breadth-first exploration is set adaptively, based on the cosine similarity between the query embedding and the entity embedding: highly relevant entities trigger deeper neighborhood exploration, while less relevant entities are explored only locally or not at all. The thresholds θa = 0.3 and θb = 0.9 are selected via an ablation study and used throughout the experiments.
Traversal respects relationship semantics: structural edges that connect chunks and documents to entities are excluded, focusing exploration on domain-relevant relationships. Retrieved information is then aggregated at three levels of granularity: chunk-level for raw textual content, entity-level for structured knowledge, and document-level for source attribution. The aggregated context is passed to a summarization LLM that produces a per-thought answer, which is added to the cumulative evidence state.
Evidence Synthesis and Source Attribution
After the iterative loop terminates, eGoT aggregates the full evidence graph, deduplicates entities, relationships, and document references, and passes the structured evidence to a synthesis LLM. The synthesis prompt is explicitly constrained to use only the provided context, and instructs the model to integrate insights across reasoning paths, resolve contradictions, address every aspect of the query, and maintain source attribution. The result is a single coherent response in which every claim is traceable to a specific source document.
Empirical Results on Standard Benchmarks
The paper benchmarks eGoT on two task families. The first is Query-Focused Summarization on the Ultradomain benchmark, evaluated across Legal, Agriculture, and Mix subsets. Following prior work, evaluation uses an LLM-as-judge protocol, scoring pairwise win rates on comprehensiveness, empowerment, diversity, and overall quality. The judge model (GPT OSS 120B) is held distinct from the models used for graph construction and retrieval. Across all three Ultradomain subsets, eGoT’s overall win rate exceeds those of HiRAG, LightRAG, NaiveRAG, GraphRAG, KAG, and a No-GoT ablation that disables the iterative thought process. The win rates against KAG, in particular, exceed 84% in every domain.
The second family of benchmarks is multi-hop question answering on HotpotQA and MultiHopRAG. On HotpotQA, eGoT achieves an exact match score of 63.0, the highest reported among the methods compared, with a competitive F1 of 71.11. On MultiHopRAG, eGoT achieves precision, recall, F1, and accuracy of 75.5%, 79.2%, 76.3%, and 79.1%, respectively, exceeding all reported baselines including TH-RAG, HyperGraphRAG, GraphRAG, LightRAG, PathRAG, and Naive RAG.
An ablation comparing eGoT against No-GoT on the Ultradomain subsets isolates the contribution of the iterative thought process: removing the graph-of-thoughts loop reduces the overall win rate by a clear margin in every domain, confirming that iterative thought generation, rather than knowledge-graph construction alone, accounts for a substantial part of the gain.
Case Study: Small Cell Lung Cancer
To probe eGoT’s behavior on a knowledge graph substantially larger than those produced by the public benchmarks, the paper constructs a knowledge graph from 1,046 full-text open-access publications on small cell lung cancer (SCLC) sourced from PubMed Central. The resulting graph contains roughly 145,000 nodes and 3 million relationships, two orders of magnitude larger than the benchmark graphs.
Twenty-one expert-curated questions of varying complexity are evaluated using LLM-as-judge metrics for faithfulness, LLM context precision, and non-vectorized context relevance. Increasing the maximum thought-expansion depth from one to three improves all three metrics: faithfulness rises from 0.80 to 0.90 (+12.4%), context precision from 0.52 to 0.66 (+27.3%), and context relevance from 0.76 to 0.92 (+21.9%). For the larger SCLC graph, in other words, the iterative reasoning loop pays off more strongly than it does on small benchmark graphs, where two iterations are typically sufficient.
Case Study: Cross-Domain Hypothesis Generation for Lupus and UV
The second case study is the most striking. UV exposure is a known trigger for lupus, with 70 to 90 percent of patients reporting photosensitivity. The full mechanistic chain that connects atmospheric composition to UV radiation, DNA damage, immune activation, and autoimmune manifestation crosses several disciplinary boundaries, including climate science, photobiology, immunology, and clinical medicine. No single corpus contains the answer.
The paper constructs a knowledge graph from two domain-specific subcorpora: 100 papers on lupus pathophysiology, photosensitivity mechanisms, and immune response, and 20 papers on UV radiation patterns, climate modeling, and atmospheric chemistry. Community-based relationship inference is used to bridge the two subgraphs. The resulting unified graph contains 31,028 entities and 598,220 relationships spanning all four domains.
For the query “How is UV related to Lupus?”, eGoT recovers a four-hop reasoning chain: UV exposure leads to keratinocyte apoptosis, which exposes autoantigens, which drive immune complex formation, which produces systemic inflammation. For the more open-ended query “Can UV forecasting affect SLE management?”, the system synthesizes evidence across climate models, photobiology, and clinical outcomes to propose three concrete hypotheses, each grounded in retrieved literature rather than parametric model knowledge. When tested on out-of-context papers, eGoT returns “no information available” rather than hallucinated content, in contrast to the baseline LLM, which produces plausible but unverifiable generic responses. Quantitatively, eGoT’s faithfulness, context precision, and source-attribution scores all exceed those of the no-retrieval baseline by wide margins (faithfulness 0.89 versus 0.52, context precision 0.71 versus 0.38, source attribution 100% versus 0%).
Limitations and Future Work
The paper is candid about open challenges. The entity standardization pipeline can over-merge near-synonymous entities (for example, “lupus nephritis” and “lupus”), and integrating biomedical ontologies as canonical grounding is identified as a natural next step. Graph quality is bounded by the extraction LLM, with missed relations, hallucinated predicates, and incorrect entity boundaries propagating into the graph; overlapping chunking and transitive and lexical inference partially mitigate this. Resolving contradictory evidence remains unsolved, since simple frequency heuristics cannot distinguish evolving knowledge from outdated or incorrect statements. Finally, while the LLM-as-judge protocol enables scalable comparison, the paper notes that human evaluation by biomedical experts is a concrete next step planned for the journal version.
Code and Paper
The full paper is available at github.com/NNeuralDynamics/eGOT, where the code and supporting prompts are also released. The paper has been accepted at ISMB 2026 and will appear in Bioinformatics.
