The Neural Dynamics Group, within The Institute for Experiential AI at Northeastern University, develops machine learning methods to address open problems across the life sciences and fundamental research. Our work spans genomics and RNA biology, protein science and engineering, biomedical applications including immunotherapy and drug synergy prediction, and core machine learning research in areas such as generative modeling and computer vision. The group is led by Ayan Paul, PhD, a Research Associate Professor with appointments at the Khoury College of Computer Sciences and affiliated with the Institute of Cognitive and Brain Health. His research background bridges theoretical physics, computational socioeconomics, epidemiology, and computational biology and biomedicine.
News & Events
Conferences, workshops, and milestones from the Neural Dynamics Group.
From the Blog
Stories from our research, our team, and our community.
-
Deep Fusion of Temporal EEG and Spatial fMRI for Cross-Subject Sleep Stage Classification
Jahnavi Umesh presents new work on graph-based multimodal fusion at the Institute for Cognitive and Brain Health, Northeastern University, demonstrating that EEG and fMRI carry complementary information for sleep stage decoding.
-
Daniela Gonzalez to Address Northeastern's 2026 Boston Graduate Commencement
A celebration of scholarship, scientific impact, and a remarkable journey from the bench to the model. Neural Dynamics is proud to see Daniela take the stage at Fenway Park.
-
Two Neural Dynamics Researchers Named to the Laurel and Scroll 100
Nihar Sanda and Daniela Gonzalez join Northeastern’s 2026 class of top graduate students, honored for research impact spanning knowledge aggregation in biomedicine and deep learning for precision oncology.
-
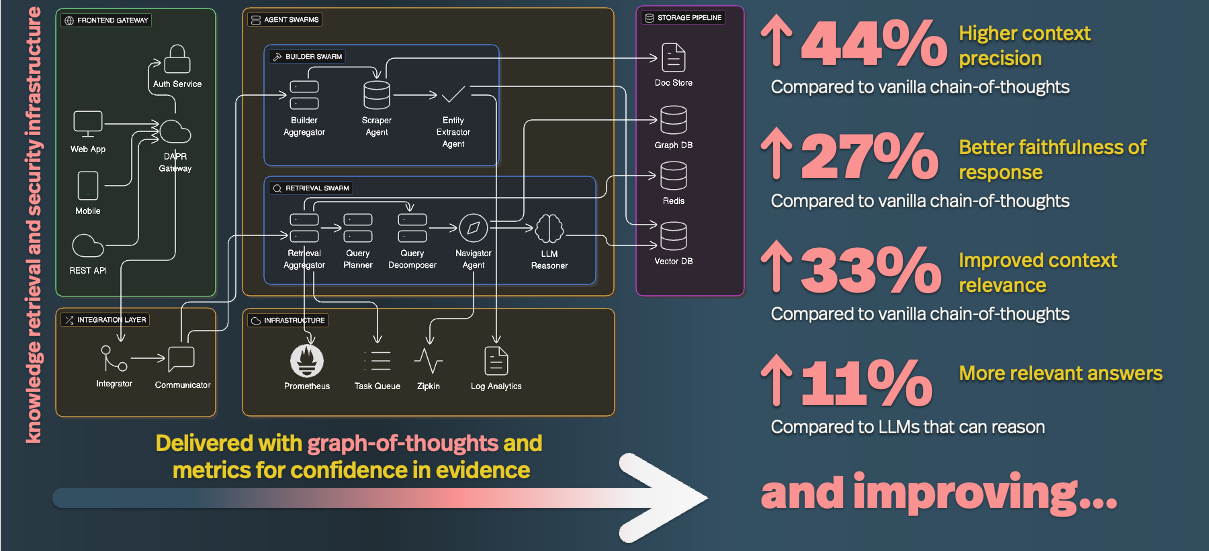
eGoT: Enhanced Graph-of-Thoughts for Multi-Hop Knowledge Retrieval and Hypothesis Generation in Biomedicine
An inside look at the lab’s ISMB 2026 paper on a retrieval-augmented generation algorithm that builds knowledge graphs directly from biomedical literature and reasons over them with an iterative graph-of-thoughts process. To appear in Bioinformatics.
-
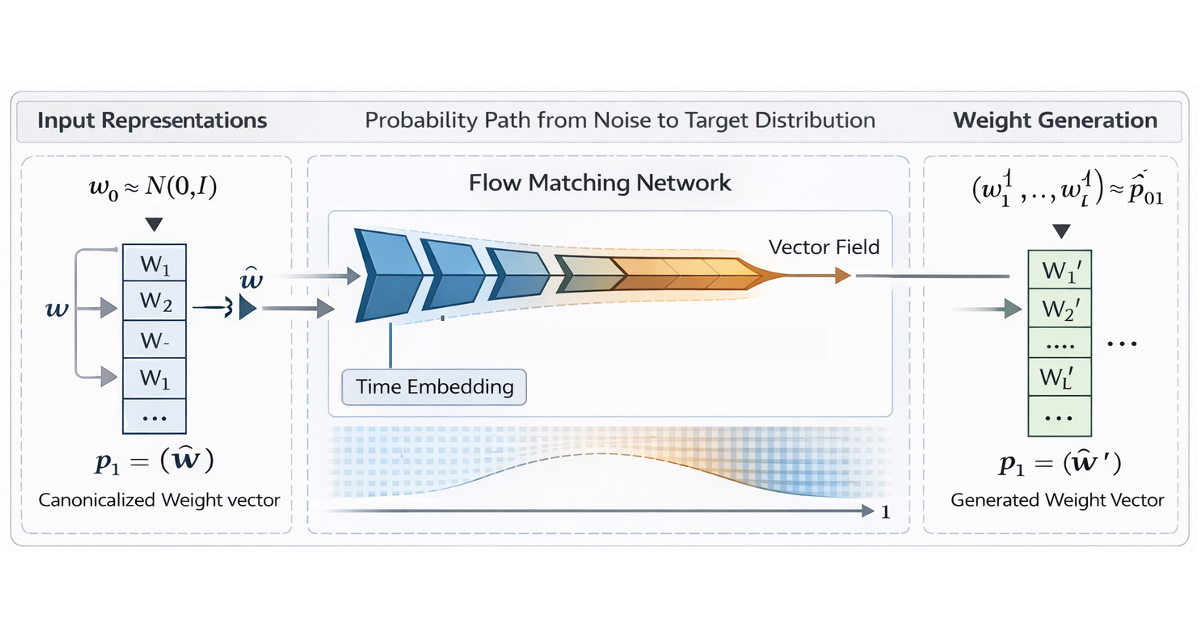
DeepWeightFlow: Re-Basined Flow Matching for Generating Neural Network Weights
An inside look at the lab’s ICLR 2026 paper on a generative model that produces complete weight sets for trained neural networks without fine-tuning, across architectures and data modalities.
Research Focus
We combine theory, computation, and data-driven methods to advance machine learning and the fundamental sciences.
Genomics & RNA Biology
Developing machine learning approaches for mRNA biology, long-read RNA sequencing, and single-cell omics to decode gene regulation in complex biological systems.
Learn more →Protein Dynamics, Function & Evolution
Applying generative models and computational methods to understand protein function, evolution, and dynamics, and to design novel proteins.
Learn more →Applied AI for Biomedicine
Leveraging ML for personalized immunotherapy, cancer subtyping, drug synergy prediction, healthcare outcomes, EEG/VEP foundation models, and epidemiological modeling.
Learn more →Core Machine Learning & Computer Vision
Advancing foundational ML through flow matching models, sensor data fusion, and image super-resolution and denoising.
Learn more →Knowledge Aggregation & Hypothesis Generation
Building LLM-powered retrieval-augmented generation systems to synthesize scientific literature, aggregate domain knowledge, and accelerate hypothesis generation across research disciplines.
Learn more →